Fine-tuning protein language model with Huggingface (Part 1)
What is this post about?
This post walks through how to fine-tune protein language models using Hugging Face libraries. It’s a polished write-up of my own learning process. When I started this, I had a good understanding of protein language models but had mainly used them to generate embeddings for downstream tasks rather than training or fine-tuning them. I was also new to using Hugging Face. I hope this guide will be useful not only for people with a background similar to mine, but also for those coming from different perspectives.
What it covers
- What it means to fine-tune a protein language model and why you might want to do it (Part 1)
- What Huggingface is and how it simplifies working with pretrained transformer models (Part 1)
- Workflow and code examples of fine-tuning pLM with Huggingface Transformer library (Part 2)
- Parameter-efficient fine-tuning with Hugginface PEFT library (Part 2)
- Actual training examples (Part 3)
Motivation: Why fine tune a protein language model?
Foundation models are becoming increasingly popular in biology across various domains. Models like ESM series and Geneformer have been trained on large-scale protein sequences and single-cell transcriptomics. Google Deepmind recnetly released Tx-LLM, a large-language model based on PaLM2 that can predict properties across modalities (small molecules, proteins, nucleic acids, cell lines, diseases). In March, they followed up with an open-source release of TxGemma, a smaller scale version of Tx-LLM.
Protein language models (pLMs) like ESM pretrained on large sequence databases learn to map protein sequence to a vector embedding in a high-dimensional representation space. The embeddings capture the proteins’ structure, biophysical properties, and evolutionary context; different characteristics may map onto low-dimensional subspaces within the embedding. Thus, it becomes quantifiable how two protein sequences are similar along certain manifolds in the high-dimensional space while different in other dimensions.
Therefore, these embeddings are very useful as input features for downstream property-prediction models. Suppose a supervised property-prediction model is trained on only hundreds of labeled sequences, which may seem too small for the model to generalize over the immense space of protein sequence and structure. If, however, the target property is mainly correlated to specific embedding dimensions and these labeled data adequately covers the distribution of natural proteins along those dimensions, the model could then accurately predict the property on novel sequences by examining the embedding values along those key dimensions.
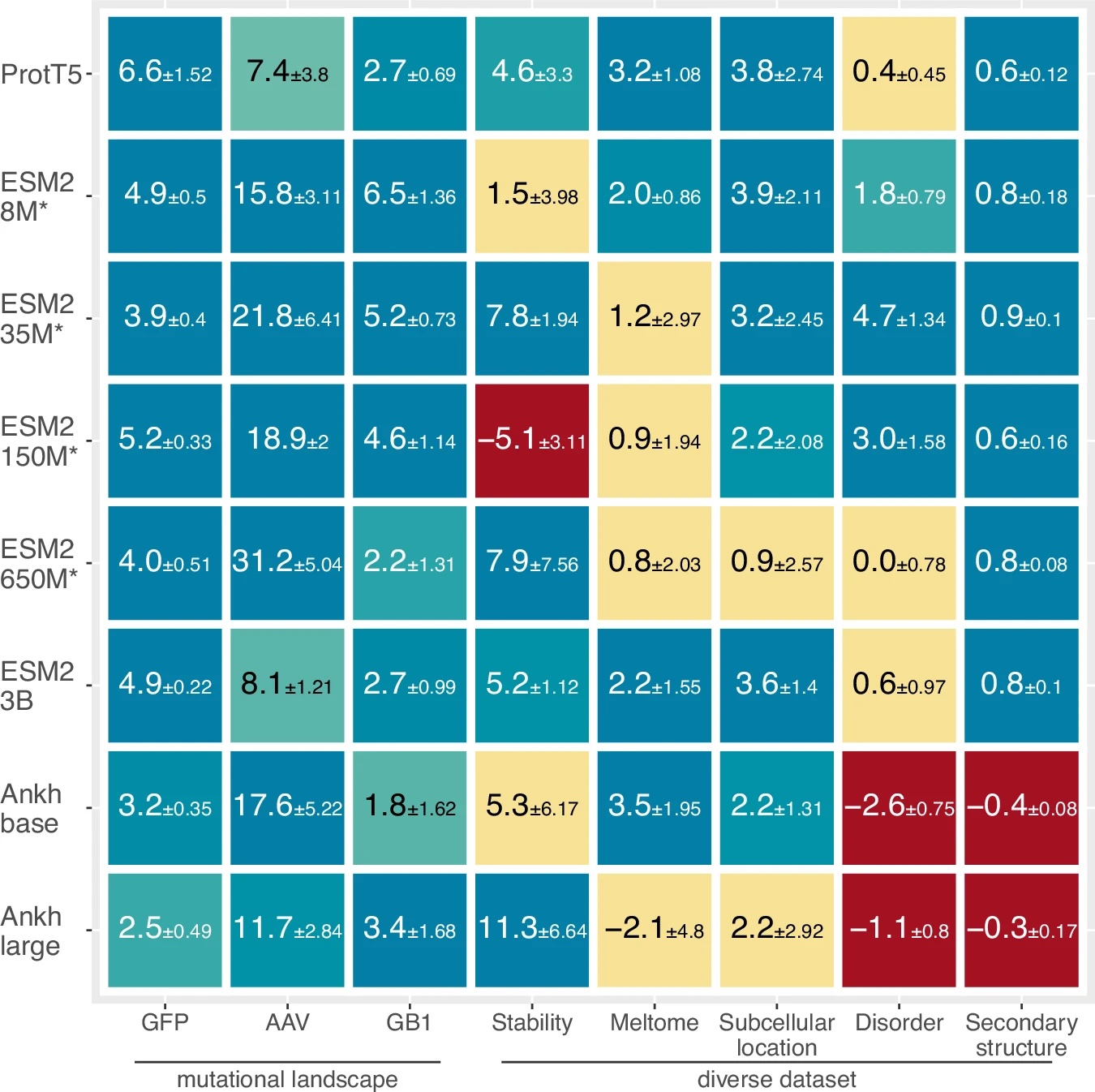
If pLMs already learn to generate information-rich embeddings from pretraining on billions of sequences, why fine-tune the pLMs to a specific task? How could this help, and couldn’t it lead to overfitting and deterioriation of generizability? I recently came across a simple and elegant paper by Burkhard Rost Lab that tested whether fine-tuning of pLMs coupled with prediction heads help improve the prediction performance. Using three pLMs (ESM2, ProtT5, Ankh) on eight different prediction tasks including predicting protein mutational landscapes, stability, intrinsically disordered region, sub-cellular location, and secondary structure, they showed nearly all of the model-task combinations improved with fine-tuning of the foundation model weights!
What’s the physical intuition for why fine-tuning helps? When we backpropagate through both the prediction head and the pLM during supervised training with labeled data, two things happen at once. First, the prediction head learns which embedding dimensions map to the target property. Second, the encoder is refined so the sequences are represented in a way that projects more cleanly onto the most relevant dimensions to the target property. It’s generally not obvious how much extra gain we get from updating the pLM itself versus just training a head on frozen embeddings, and requires empirical testing. In principle, however, this joint adjustment can yield a representation that’s better aligned with the prediction task.
Fine-tuning vs. Domain-adaptive pretraining
The term “fine-tuning” is used in at least two separate contexts in literature.
- Scenario 1: A prediction task uses a pLM embedding as input and outputs property value. Supervised training of this prediction head backpropagates through both models. For example, for predicting the fitness of GFP protein mutants in Schmirler et al., the sequences are embedded by ESM2 and then passed into a MLP classifier head. During the supervised training of the MLP head, the loss on mutational effect prediction is used to update the weights of both the MLP head and the ESM2 model.
- Scenario 2: This scenario takes a pLM trained on wide corpus of sequences, and continues to train with the same objective but with a more specific dataset. For example in Madani et al, ProGen model is first trained on 281 million protein sequences from UniParc, UniProtKB, Pfam, and NCBI. ProGen is a generative model and is trained with next-token prediction objective. Then, before it’s used for generating artificial lysozyme sequences, it’s fine-tuned with 55,948-sequences that belong to phage lysozyme (PF00959), pesticin (PF16754), glucosaminidase (PF01832), glycoside hydrolase family 108 (PF05838) and transglycosylase (PF06737) from the Pfam family. Notably, in this case the objective function for fine-tuning is still the next-token prediction loss.
These scenarios show that the term fine-tuning can be used broadly in the field. Here are more precise definitions for distinguishing them:
- Task-adaptive fine-tuning (TaFT)
- What it is: Pretrained foundation model is attached to a downstream prediction task model, and the weights are updated using a supervised loss for the downstream property-prediction task (i.e. Scenario 1)
- When to use it: When there are enough labeled examples of the target property that we can learn how to shift the foundation representations themselves to better separate the examples.
- Domain-adaptive pretraining (DaPT)
- What it is: Pretrained foundation model continues to be trained on its original self-supervised objective (e.g. masked language modeling) but using a small, specialized dataset (i.e. Scenario 2)
- When to use it: When the target proteins are from a niche family whose statistics differ substantially from the base model’s training set. The idea is to realign the foundation’s language to the domain of interest before any supervised step.
The key distinction is that domain-adaptive pre-training simply consists of training the model on its original objective. In contrast, task-adaptive fine-tuning requires connecting a foundation model with a prediction task head and backpropagating through both. For the rest of this post, we will focus on task-adaptive fine-tuning.
Parameter-efficient fine-tuning
Before diving into the implementation of fine-tuning, let’s consider its two main downsides: cost and risk.
- Cost: Full fine-tuning of a foundation model with hundreds of millions to tens of billions of parameters can be expensive and complicated. Training very large models will require multiple GPUs with distributed training and complex checkpointing workflows.
- Risk: Updating pLM weights based on limited supervised tasks opens the door to overfitting. pLM pretraining already results in embeddings that contain rich evolutionary, structural, and physicochemical information. After the fine-tuning updates the model may end up forgetting these fundamental information and instead memorizing the idiosyncrasies of the small labeled training set. This phenomenon of losing the previously learned knowledge after fine-tuning is called catastrophic forgetting.
A strategy that can mitigate both of these problems is parameter efficient fine-tuning (PEFT). Common PEFT approaches include:
- Partial fine-tuning
- Weights from only certain parts (e.g. the final transformer block) are updated, while the others are frozen
- Adapters
- Additional layers are inserted within the model (rather than on top like the prediction head) and only these are trained, while the original model weights are frozen
- Reparameterization (Low Rank Adaptation; LoRA)
- Updates to weight matrices are allowed only as a product of two lower rank matrices, drastically reducing the number of trainable parameters
There are many other resources to learn more about the methods of PEFT. This blog post from IBM is a nice introduction. In this post, I’ll mainly look at the implementation of LoRA.
Implementation of task-adaptive fine-tuning with Huggingface
Challenges of using pretrained pLMs
As noted before, task-adaptive fine-tuning requires connecting a pretrained pLM with a prediction task head and backpropagating through both. Let’s think of the challenges as we try to implement this. Because it’s hard to predict which pLM may generate the best embedding for our downstream task (see the above figure from Schmirler et al: performance of different pLMs vary for downstream tasks), we want to test several pLMs. We will face the following time-consuming and tedious tasks to get started with the various pLMs:
- Installing multiple pLM packages or cloning the repos
- Creating and managing environments for each model
- Reading through documentations and figuring out model-specific quirks, such as handling special tokens or understanding the correct arguments for the
forward()method - Reading through the code repos to understand the quirks if documentation isn’t great
Adding prediction head and fine-tuning will create more headaches, such as:
- Having to subclass the pretrained pLM and attaching the task head
- Resolving odd dependency issues that arise
- Implementing PEFT by directly modifying the pLM architecture
Standardization of existing transformer models
Fortunately, these problems boil down to mainly three common issues:
- Setting up the model: having to install/clone, with full environment or container setup for each model
- Using model-specific syntax: tokenizer, attention mask, padding, etc.
- Wiring with prediction head with the pLM and fine tuning: required for fine-tuning with supervised task
Given that these are structured, repeated problems, people have developed a framework that can be used to remove much of the pain from them! That is what Hugginface does.
Huggingface helps solve these issues by providing a unified interface to access and subclass various pretrained transformer models. Through the transformer library we can use most of existing pLMs through a unified syntax. Let’s quickly see how it helps with each of the three pain points before we move onto practical implementation of fine-tuning in the next post.
- Setting up the model → Import from Huggingface transformer library
- Huggingface provides a single, simple API (
AutoModel.from_pretrained) to load pretrained models and tokenizers from the Huggingface Hub, automatically handling the model architectures and pretrained weights. Thus, there is no need to install or clone each model from its repo. For example, generating ESM2 embedding for proteins looks like this.
from transformers import AutoModel, AutoTokenizer # These libraries handle loading tokenizer and model from name # Model and sequence examples model_path = "facebook/esm2_t33_650M_UR50D" sequences = [ "MKTAYIAKQRQISFVKSHFSRQDILDLIC", "GQDPYEEIVIAFINKPRLQYF", "VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFP" ] # Load ESM2 tokenizer and pretrained 650M-parameter model tokenizer = AutoTokenizer.from_pretrained(model_path) model = AutoModel.from_pretrained(model_path) # Generate embedding inputs = tokenizer(sequences, return_tensors="pt") #Returns BatchEncoding object with input_ids and attention_mask outputs = model(**inputs) #Returns ModelOutput class with loss, logits, hidden_states, attentions embeddings = outputs.last_hidden_state # shape: [batch_size, sequence_length, hidden_dim]If we want to use a different model, ProBert, we need to change the model path:
model_path = "Rostlab/prot_bert"But we also need to do one more thing. The tokenizer for ProBert expects sequences to be spaced, e.g. “M K T A” rather than “MKTA”. Thus we must add
sequences = [' '.join(list(sequence)) for sequence in sequences]before passing it to the tokenizer.
This second example shows that while Huggingface abstracts away many of model-specific elements, we still need to understand some quirks about the models we want to use. Also, Huggingface does not directly handle separate environments or conflicting dependency versions for each model. So if there’s a clash in libraries required by two models, this will require careful virtual environment setup. However, as long as common dependency versions for packages are compatible across multiple models, which is often the case, then multiple transformer models can be used in the same environment without needing separate isolated setups.
- Huggingface provides a single, simple API (
- Using model-specific syntax → Using unified syntax
- Huggingface standardizes input/output data structures (tokenized inputs, attention masks, positional encodings and
ModelOutputobjects), so methods like.encode(),.decode(),.forward(), and.generate()work the same way across different transformer models.
- Huggingface standardizes input/output data structures (tokenized inputs, attention masks, positional encodings and
- Wiring with prediction head and fine-tuning → Simplified by unified API
- Huggingface provides built-in utilities (
Trainer,TrainingArguments) that standardize training loops, logging, evaluation, hyperparameter tuning, and distributed training. - The
PEFT(for parameter-efficient fine-tuning) library provides ways to implement fine-tuning techniques like LoRA with any model from thePreTrainedModelclass.
- Huggingface provides built-in utilities (
These second and third points will become clearer when we look at practical implementation of fine-tuning pLMs.
Workflow for fine-tuning
Before getting into the code, let’s conceptually break down what a full parameter pLM fine-tuning for a prediction task requires.
- Define the prediction head (classification/regression) that uses pLM embedding as input.
- Define the main model that wires the pLM and the prediction head together. Initialize the pLM with pretrained weights.
- Prepare labeled datasets for supervised training, validation, and test.
- Define optimizer and trainer.
- Pipeline to bring everything together.
We will see that PEFT implementation just requires one additional simple step. Also along the way, we will need to pay attention to correctly handle tokenization, attention mask, and padding/truncation. But Huggingface framework will help us with these too.
Setting the goals for what the code should do
In Part 2, I will share some code examples that correspond to the steps outlined above. But before that, it is important to note that when developing a model, we may want to test out multipel pLMs in combinations with multiple task head architectures. Therefore, it can be hugely beneficial to have a nice modular code that:
- can be used with various pLMs as plug-and-play
- provides some template for simple prediction head, but can also work with other custom prediction heads by allowing passing of additional arguments.
Although huggingface provides a convenient interface, there are still some model-specific quirks that made doing this somewhat tricky. For example, to enable various prediction heads to work plug-and-play, the following issues must be considered.
- Models may make residue(token)-level or protein-level prediction, and may do classification or regression. We need to use appropriate loss function for each case. To break down each case:
- Residue-level Classification
- Example: does each residue belong to intrinsically disordered region?
- Loss function: cross entropy loss summed over residues (excluding padded residues)
- Residue-level Regression
- Example: per-residue evolutionary mutability
- Loss function: MSE loss, MAE loss, etc. summed over residues (excluding padded residues)
- Protein-level Classification
- Example: classify a protein as binder or non-binder to a given target
- Loss: cross entropy loss on protein sequence-level logits
- Protein-level Regression
- Example: prediction of melting temperature Tm
- Loss: MSE loss, MAE loss, etc at protein sequence level
- Residue-level Classification
- For making a protein-level prediction, there are different ways of aggregating the embeddings across the residues. For example, some BERT-based models like ProtBert can have the special
clstoken that can be used for classification. User may choose to use it, or ignore that and take the mean of the embeddings across the residues.
Moreover, the pLM themselves will also have model-specific quirks:
- Data pre-processing steps needs to be correctly handled. As pointed out earlier, the ProtBert model requires uppercase amino acids that are separated by spaces. Other mothers may have their own quirks.
- Each pLM will have certain model-specific attributes in the model architecture. For example, T5-based models like
ProtT5hasself.sharedlayer that implements vocabulary encoding. The namesharedcomes from the fact that it is a shared layer by the encoder and decoder. Naturally, encoder-only models likeESM2will not have this layer. If we want a modular class for our main model that enables plug-and-play with different pLMs, we should avoid referencing specific attributes like this and only use attributes that are universal for thePreTrainedModelclass in transformer library.
When we look at RSchmirler et al. repo, they defined separate classes for fine-tuning different pLM models (e.g. T5EncoderForSimpleSequenceClassification and load_T5_model for ProtT5, load_esm_model for ESM2, etc) to handle different model quirks. Similarly, different tasks are handled by their own classes (e.g. T5EncoderForTokenClassificaion and T5EncoderForSimpleSequenceClassification are defined separately although most of the functionality are same). While this works for the scope of their study, it would be nice to have a more modular framework.
Next Steps
In Part 2, I will go through code examples that implements the steps mentioned above for pLM fine-tuning.